Protocol Number 1: Bacterial Transformation
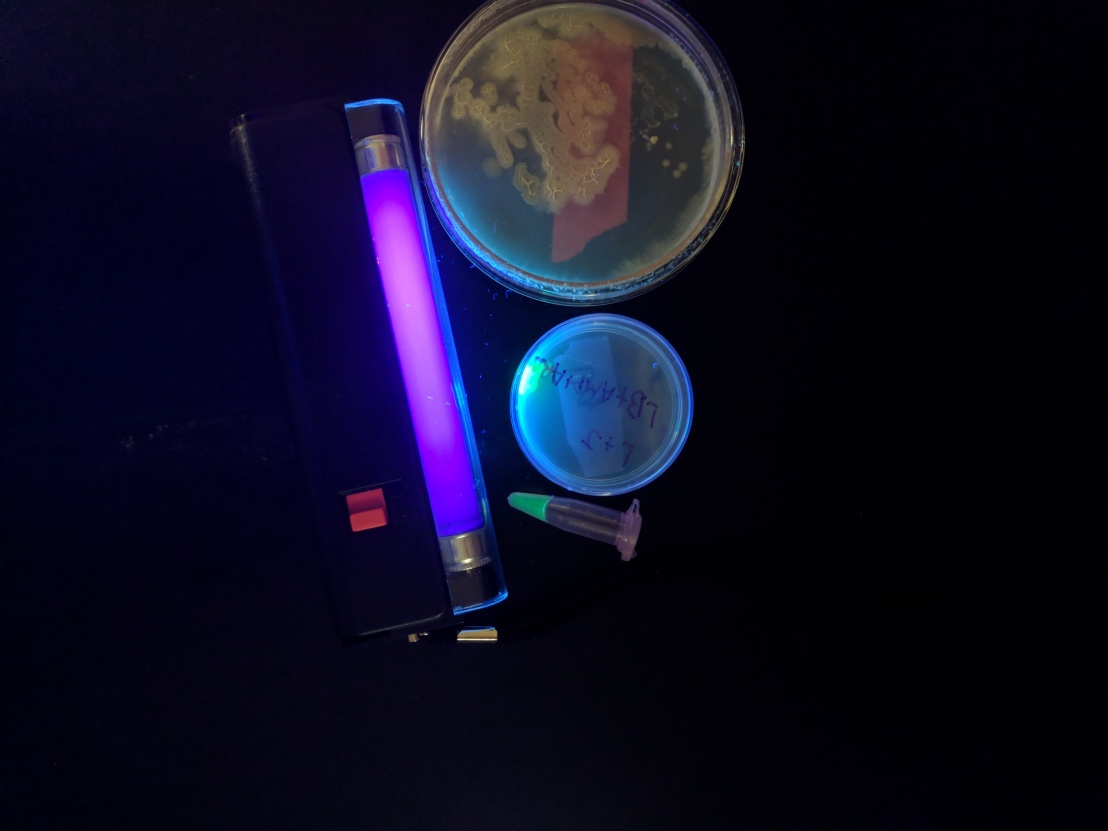
On 10/20/17 my lab partner and I performed a bacterial transformation lab on E.Coli to express a Green Fluorescent Protein (GFP). This process is done by using a plasmid that codes for the GFP and inserting the plasmid by using transformation solution and heat shock. The transformation solution coats the DNA with a non-polar sheath to allow it to interact with the cell membrane of the E.Coli to allow it to pass through the phospholipid bilayer membrane of the cell which consists of mainly non polar regions. Combined with heat shock which allows the cell wall to momentarily break open, allows the most amount of uptake of plasmid. Then allowing the E.Coli to incubate allows satellite colonies to grow and express the GFP.

The picture above shows the E. Coli with the one satellite colony expressing the GFP. This result is not what we were expecting because we expected multiple colonies to grow and express GFP. We figure that most of the error came from aseptic technique. I say the because if the error lied in our ability to do the procedure we would not have seen any data.
Protocol Number 2: Inoculation of Liquid Media with Transformed Cells

In order to to complete the isolation of the GFP the bacteria needs to be grown in a liquid media. The liquid media is exactly like the agar plates however when making the liquid medium it lacks the protein that would normally solidify the agar.
This protocol was very straight forward in the sense that the only work to be done was to transfer the E.Coli colony from the plate to the liquid media and allow it to incubate over night with occasional breaks where the tubes need to be shook to increase the levels of oxygen.
Protocol Number 3: Concentration of Bacteria
In this step of the eventual isolation of GFP, the main goal of this is to break open the cell wall and allow everything inside the cell to begin spilling out. This is done by taking the liquid media that has incubated the E.Coli and transfer a certain amount into a micro centrifuge tube. At this point the tubes are centrifuged at a high velocity for about five minutes. This allows the bacteria to move its was towards the bottom of the tube. This can be seen because after the 5 minutes are up a glowing green pellet can be seen at the bottom of the tube. It is at this point the liquid media is removed and a buffer is added to re-suspend the pellet to get ready for the break down of the cell wall. A drop of lysozyme is added which will initiate the break down of the cell and with that done the tubess need to be kept at a very cold temperature because freezing the tube will also aid in the process of breaking open the cell.

Protocol Number 4: Bacterial Lysis
In this part of the lab the main objective is to further break open the cell wall and prepare the HIC column. The reason why ensuring the cell wall is completely destroyed is because it allows for the most amount of GFP to be collected at the end. This is because if GFP if still inside the cell wall and we “filter” using the HIC column not all of the GFP will make it to the bottom to be collected. And on top of ensuring the cell wall is open the HIC column needs to go through a “pre-run” much like a gel in gel electrophoresis needs to have a quick 2 min pre run.

Protocol Number 5: Hydrophobic Interaction Chromatography (HIC)
It is in this step of the protocol where the GFP and other gunk can be filtered and only leave the GFP protein to be collected. The way this works is that a series of buffers are are added to HIC column. What each buffer does is bind to different particles in the solution making it harder and harder from them to mover throughout the column. What this will eventually do for us is have all the gunk stick to white substance that is in the column while the GFP is allowed to move move much more easier that the other particles. In the end of this run there will be a little less that 1ml of pure GFP collected in a tube. The separate test tubes are conserved to do later experiments with (pertaining to concentration of a solution).

My Thoughts
After completing this lab my brain was completely blown away at the idea of being able to collect a specific protein from any bacteria you want. The skills required to do this was shockingly doable and requires some knowledge of chemistry because when it gets to the point where you filter knowing how the chemistry of each particle interacts with one another is important to know. Overall I was shock at the success we had and opens my mind to all the possibilities and implications for iGEM.